‘The Joy of the Discovery’: An Interview with Jennifer Doudna
11 de juny 2020
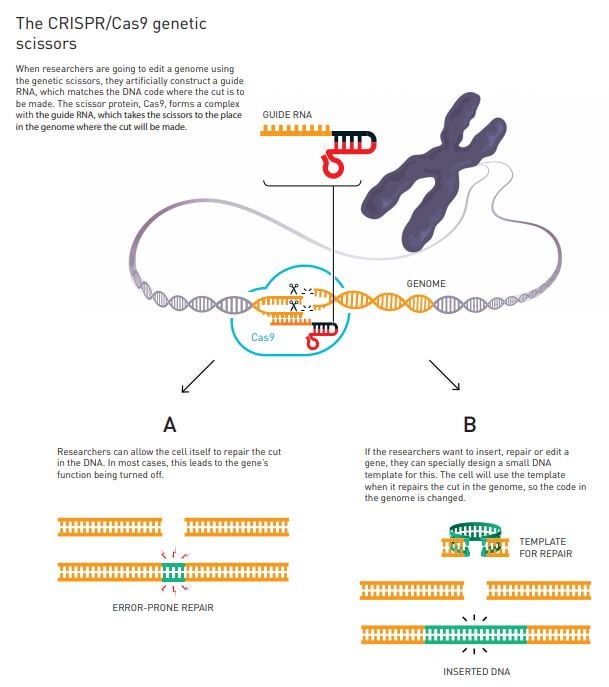
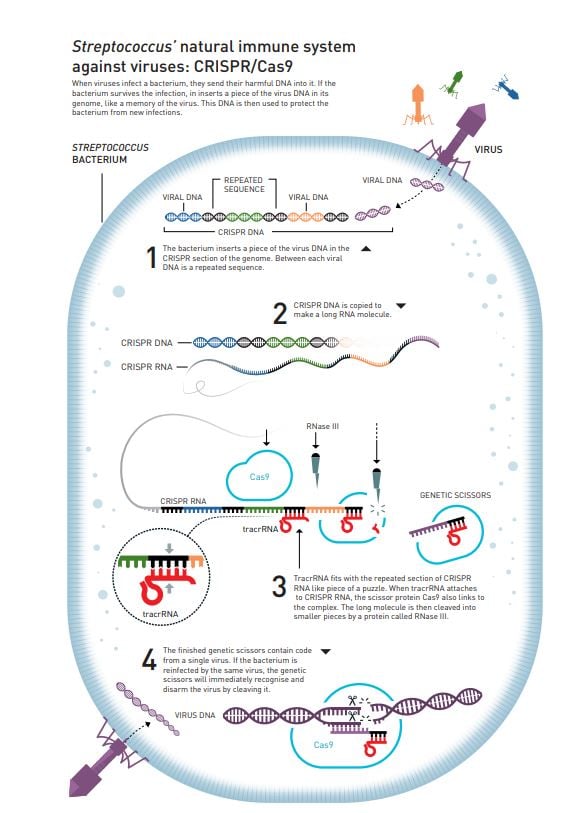
Why CRISPR is a breakthrough
‘The Joy of the Discovery’: An Interview with Jennifer Doudna
03 d’agost 2017
A quantum leap in biomedicine
Today you should have a look at Nature. The reason is genome editing of human embryos. Although a chinese team performed something similar in 2015, right now the details are clearly explained. The trial has been developed to fix a mutation that causes hypertrophic cardiomyopathy.If you don't want to read the whole article, check the news. Genome editing is gaining momentum as I said some months ago, it is closer than you may think.
PS. Why FBI is concerned about CRISPR? The answer in FT Big Read
PS. The news in FT.
PS. A useful article from Nature 2015:
Genome editing: 7 facts about a revolutionary technology
Article toolsediting is in the spotlight again as a large international meeting on the topic is poised to kick off in Washington DC. Ahead of the summit, which is being jointly organized by the US National Academy of Sciences, the US National Academy of Medicine, the Chinese Academy of Sciences and Britain’s Royal Society and held on 1–3 December, we bring you seven key genome-editing facts.
07 d’octubre 2020
CRISPR Nobel prize
Genetic scissors: a tool for rewriting the code of life
The Royal Swedish Academy of Sciences has decided to award the Nobel Prize in Chemistry 2020 to
Emmanuelle Charpentier, Max Planck Unit for the Science of Pathogens, Berlin, Germany
Jennifer A. Doudna, University of California, Berkeley, USA
“for the development of a method for genome editing”
Popular information: Genetic scissors: a tool for rewriting the code of life (pdf)
Scientific Background: A tool for genome editing (pdf)
Unfortunately, the Royal Swedish Academy of Sciences has shown its ignorance about the real discovery of CRISPR. It happened in the '90s in Salines de Santa Pola by Dr. Martinez Mojica.
08 de setembre 2017
The long and bumpy road to CRISPR
I've read the same book than Diane Coyle this summer. If you want a clear understanding of what's going on in genomic editing, it should be your first choice. A crack in creation is a description and analysis by Jeniffer Doudna the main researcher on the topic. For those that are excited by genome editingit is good to read this statement:
It’s easy to get caught up in the excitement. The fact that gene editing might be able to reverse the course of a disease—permanently—by targeting its underlying genetic cause is thrilling enough. But even more so is the fact that CRISPR can be retooled to target new sequences of DNA and, hence, new diseases. Given CRISPR’s tremendous potential, I’ve grown accustomed over the past several years to being approached by established pharmaceutical companies asking for my help in learning about the CRISPR technology and about how it might be deployed in the quest for new therapeutics.Therefore, caution is required and ethical implications are huge as I've said before.
But therapeutic gene editing is still in its infancy—indeed, clinical trials have only just begun—and there are still big questions about how things will progress from here. The decades-long struggle to make good on the promise of gene therapy should serve as a reminder that medical advances are almost always more complicated than they might seem. For CRISPR, too, the road leading from the lab to the clinic will be long and bumpy.
Deciding what types of cells to target is one of the many dilemmas confronting researchers—should they edit somatic cells (from the Greek soma, for “body”) or germ cells (from the Latin germen, for “bud” or “sprout”)? The distinction between these two classes of cells cuts to the heart of one of the most heated and vital debates in the world of medicine today.
Germ cells are any cells whose genome can be inherited by subsequent generations, and thus they make up the germline of the organism—the stream of genetic material that is passed from one generation to the next. While eggs and sperm are the most obvious germ cells in humans, the germline also encompasses the progenitors of these mature sex cells as well as stem cells from the very early stages of the developing human embryo.
Somatic cells are virtually all the other cells in an organism: heart, muscle, brain, skin, liver—any cell whose DNA cannot be transmitted to offspring.
Highly recommended.
09 de maig 2019
Genome editing: the game of biology is about to change
The foundations of gene editing came about because a scientist in Alacant, Dr. Mojica started to find weird DNA sequences in some bacteria he was studying. After that Profs. Doudna and Charpentier and later Prof. Zhang translated initial findings into practice. Therefore it all started when a microbiologist studied the arms race between bacteria and viruses.
You'll find all these details in a book by Nessa Carey. If you want to understand in plain words what CRISPR is and what may represent for biology, then you have to read it.
The gene editing revolution is creating a technological toolkit that almost any half-decent scientist can lean into and find something useful. On the one hand, that should make us very excited. We can both solve problems and simply indulge our curiosity. But should it also make us worried? Using chisels and a mallet, Michelangelo created some of the most exquisite sculptures we have ever seen. But give the same heavy, sharp tools to someone else, and we can get a very different and much bloodier outcome.
But the same technology can also be used to alleviate human suffering, and if we are smart enough, lessen the impact that our heavy-footed species has on the only planet we know of in the entire universe that supports complex life. We cannot un-invent this technology, we probably can’t even control its spread. So what choice do we really have but to embrace it and use it well, to create a safer, more equal world for all?
22 d’abril 2021
04 de novembre 2017
How to change individual letters of your DNA?
These are exciting times for genetic research, though we'll have to wait for specific clinical applications
14 de febrer 2020
Repairing DNA: a review
Jenifer Doudna publishes a must read review article on genome editing in Nature this week.
Current clinical trials using the CRISPR platform aim to improve chimeric antigen receptor (CAR) T cell effectiveness, treat sickle cell disease and other inherited blood disorders, and stop or reverse eye disease. In addition, clinical trials to use genome editing for degenerative diseases including for patients with muscular dystrophy are on the horizon.
Notably, all of the genome-editing therapeutics under development aim to treat patients through somatic cell modification. These treatments are designed to affect only the individual who receives the treatment, reflecting the traditional approach to disease mitigation. However, genome editing offers the potential to correct disease causing mutations in the germline, which would introduce genetic changes that would be passed on to future generations.
At the time of writing, international commissions convened by the World Health Organization (WHO) and by the US National Academy of Sciences and National Academy of Medicine, together with the Royal Society, are drafting detailed requirements for any potential future clinical use.Meanwhile, CRISPR is closer than you think.
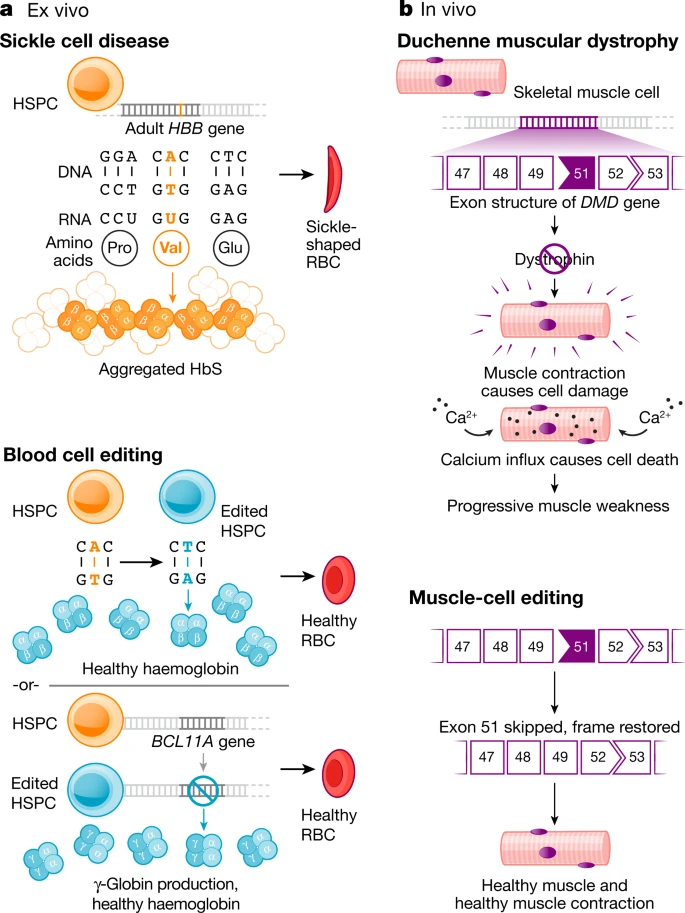
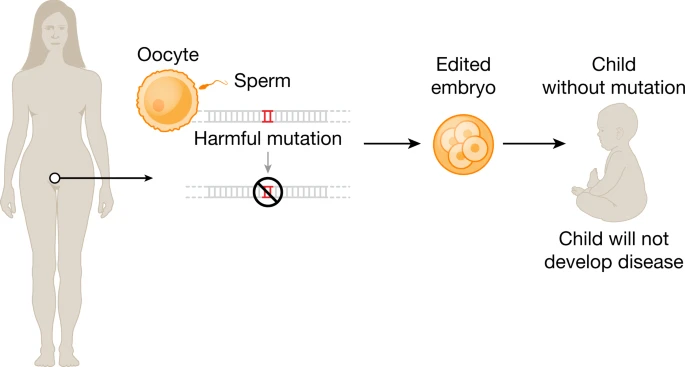
Fig. 1: Ex vivo and in vivo genome editing to treat human disease.
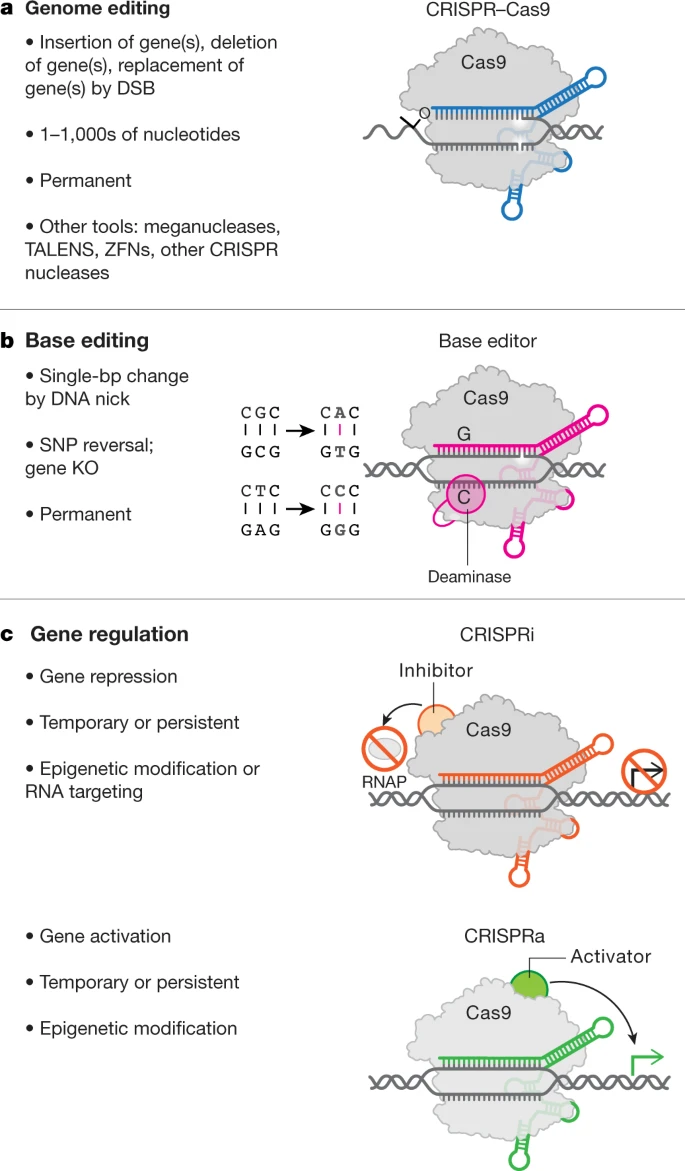
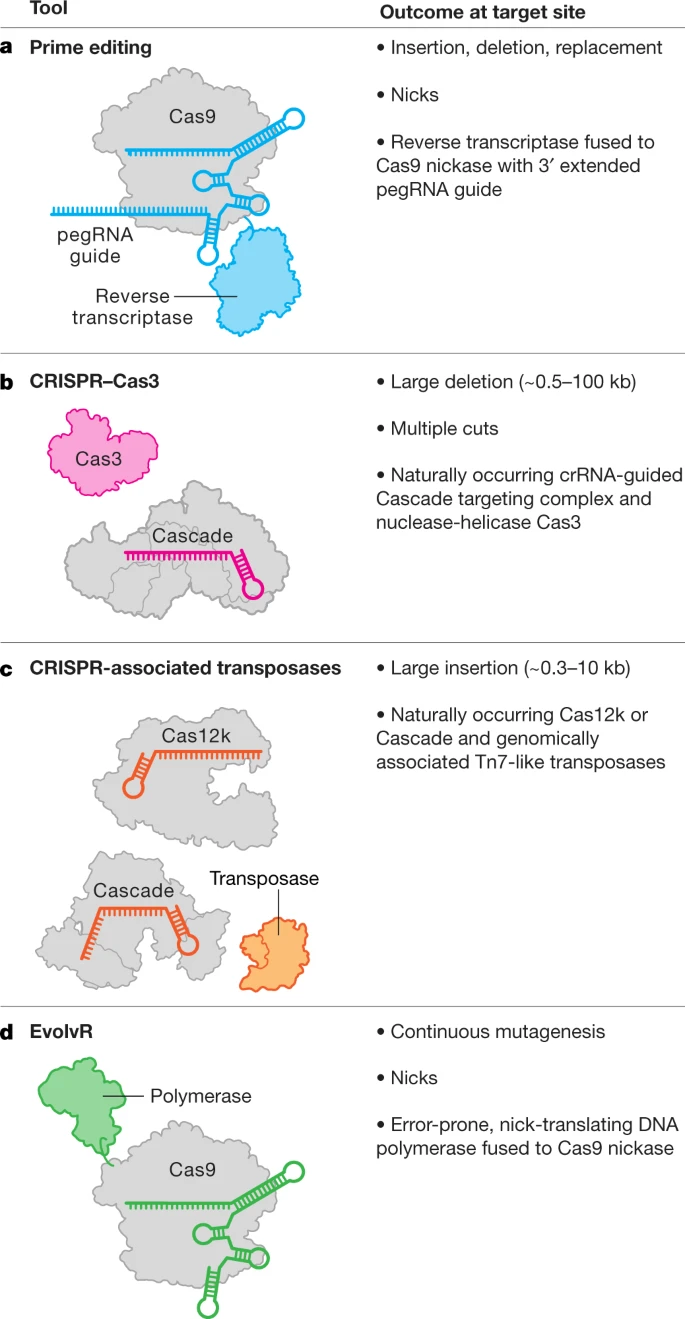
Fig. 2: The genome editing toolbox.
17 de novembre 2023
Els nirvis de la indústria farmacèutica europea
La proposta de nova regulació farmacèutica europea ha desfermat la preocupació a una indústria que ja havia d'estar preocupada abans que això passés. I és que només veient els medicaments que s'han aprovat els darrers temps i la inversió en recerca recent, sabem que anem a empentes i rodolons.
En aquest context hi ha nirvis de la indústria sobretot per dues coses: per la protecció de dades regulatòries i per l'exclusivitat de mercat dels medicaments orfes. Diguem-ho clar, perquè la regulació de patents segueixi protegint d'igual forma el monopoli temporal. Aquest és el concepte. Després ve el detall de facilitar l'accés als medicaments per part dels pacients, però això ja són els serrells.
Ahir UK ja va aprovar el Exa-cel, el medicament de CRISPR Therapeutics i ho va fer amb dues indicacions, anèmia de cèl·lules falciformes i beta-thalassemia. És la primera vegada al món que s'aprova definitivament una teràpia genètica ex-vivo basada en CRISPR. A la FDA un consell assessor va donar la seva aprovació que s'espera sigui definitiva el dia 8 de desembre.
Curiosament CRISPR Therapeutics té la seu social a Europa ,no-UE , a Suïssa, a Zug. La va fundar Emmanuelle Charpentier fa 10 anys, guanyadora del Nobel amb Jennifer Doudna. Ara bé, només hi té la seu social. Tot, tot s'ha fet junt amb Vertex a Boston. Si mirem la vacuna de Pfizer va ser originada a Alemanya (Biontech), però va comercialitzar-se com nord-americana. Podríem repassar molts més exemples on la recerca europea és potent i la comercialització s'esvaeix. Però això no ho arreglarà la protecció mitjançant patents.
La indústria europea ens explica en un document quina serà la seva estratègia de lobby per als propers mesos i després d'explicar que la redacció actual provocarà el diluvi universal (menys accés als medicaments), diu el que cal fer (RDP vol dir protecció de dades regulatòries i OME vol dir exclusivitat de mercat de medicaments orfes):
- In line with the European Council Conclusions (March 2023), Europe needs to strengthen, rather than cut, the region’s RDP baseline and OME.
- Providing meaningful and predictable incentives, attainable fairly, that would encourage additional R&D investment compared to today.
- Jointly addressing barriers and delays to access based on a shared understanding of the evidence generated by the European Access Hurdles Portal.
- Limiting Bolar exemption for activities related to seeking regulatory approval.
- Developing a patient-centred, more inclusive definition of unmet medical need.
- By acknowledging the value of innovation and encouraging advancements in prevention, treatments and care, Europe can ensure that no patient is left behind.
- A robust framework for mechanism of action Paediatric Investigation Plans (PIPs) is essential to ensure that this new obligation is effective to achieve its purpose and is manageable for developers.
- Further optimising the regulatory framework and ensuring maximum use of expedited pathways in support of patient needs.
- Ensuring that supply chain and environmental requirements are proportionate and fit-for-purpose while not prohibiting or delaying patient access to medicinal products.
14 de març 2021
CRISPR: from lab to the clinic
This is the year that CRISPR moves from lab to clinic
Jennifer Doudna says:
In 2021, researchers will use CRISPR to enhance our medical response to the Covid-19 pandemic. Teams will continue to collaborate and bring to market vital CRISPR-based diagnostic tools that are accurate, rapid and painless. One currently being developed and scaled by Mammoth Biosciences, a company I co-founded, along with partners at the University of California, San Francisco and the pharmaceutical company GSK, can detect and indicate the presence of SARS-CoV-2 RNA in a similar fashion to a pregnancy test.
CRISPR will also have an important effect on the way we treat other diseases. In 2021, we will see increased use of CRISPR-Cas enzymes to underpin a new generation of cost-effective, individualised therapies. With CRISPR enzymes, we can cut DNA at precise locations, using specifically designed proteins, and insert or delete pieces of DNA to correct mutations.
This is precisely what is going on.
19 de juliol 2023
Canviant la funció de producció de les proves diagnòstiques
Real-Time, Multiplexed SHERLOCK for in Vitro Diagnostics
Durant la pandèmia es va produir una innovació notable a la tecnologia de proves diagnòstiques. Va passar desapercebuda per alguns, però no pels que llegiu aquest blog. Es tracta d'utilitzar CRISPR que inicialment s'ha desenvolupat per a edició genètica, per a la detecció d'àcids nucleïcs en un sol pas, i per tant també de Sars-COV-2. La dificultat d'aplicació que tenia aquella prova, que va rebre el vist-i-plau de la FDA era que el procés no es podia automatitzar, i per tant calia amplificació prèvia. Ara acaba de publicar-se la prova definitiva que pot capgirar moltes coses, i que per tant pot canviar la funció de producció dels laboratoris. La troballa d'enzims termoestables ha estat la qüestió clau per a l'èxit de l'equip de Feng Zhang.
Cal dir que la companyia rival, Mammoth Biosciences, de Jennifer Doudna, també té en marxa una prova similar: DETECTR BOOST.
I ara ja ha començat el procés de patentar enzims. Tant per una empresa com per l'altre. I això esdevé inadmisible si el que es pretén patentar la natura o variacions sobre la natura. Però ningú se n'està adonant , als USA ja s'ha fet tard, però a Europa encara hi som a temps.
N'estic segur que les patents, una vegada més, limitaran l'accés a aquesta gran innovació.
Cartier-Bresson
PS. Per si voleu saber com funciona DETECTR BOOST
PS. Per tal de comprendre l'abast del que signifiquen les proves diagnòstiques basades en CRISPR el millor és consultar un article de revisió com aquest. I la taula següent conté les dades bàsiques:
Characteristics of reported CRISPR-based diagnostics
From: CRISPR-based diagnostics
Name | Enzyme | Preamplification | Assay timea (min) | Sample preparation | Readout | Applications | LODc (mol l−1) | LODc (copies per ml)3 | References |
|---|---|---|---|---|---|---|---|---|---|
CRISPR type II | |||||||||
NASBACCb | Cas9 | NASBA | 120–360 (one pot) | Column-based or crude extraction | Colometry | Discrimination between African and American ZIKV | 1.0 × 10–15 | 6.0 × 105 | |
CRISPR–Chip | Cas9 | – | 15 | Column-based | Electrochemical | Detection of gDNA from cell lines and DMD patients | 2.3 × 10–15 | 1.4 × 106 | |
CRISDA | Cas9 nickase | SDA | 90 | Column-based | Fluorescence | Detection of gDNA; breast-cancer-associated SNPs in cell lines | 2.5 × 10–19 | 1.5 × 102 | |
FLASH | Cas9 | PCR | NS | Column-based | NGS | Detection of gDNA; antimicrobial resistance genes in clinical samples | 1.9 × 10–18 | 1.1 × 103 | |
CAS-EXPAR | Cas9 | EXPAR | 60 | Chemical (phase separation) | Fluorescence | Sensing of methylated DNA; L. monocytogenes mRNA | 8.2 × 10–19 | 4.9 × 102 | |
Cas9nAR | Cas9 nickase | Strand-displacing DNA polymerase | 60 | Column-based | Fluorescence | Detection of bacteria (S. typhimurium, E. coli, M. smegmatis, S. erythraea); detection of KRAS SNPs in cell lines | 1.7 × 10–19 | 1.0 × 102 | |
CRISPR type V | |||||||||
DETECTR | Cas12a | RPA | 10 (RPA) and 60–120 (CRISPR) | Crude extraction | Fluorescence | Detection of HPV16 and HPV18 in human samples | 1.0 × 10–18 | 6.0 × 102 | |
Cas14-DETECTR | Cas14 (Cas12f) | PCR | NS (PCR) and 120 (CRISPR) | Crude extraction | Fluorescence | Detection of HERC2 SNPs in human samples | n.s. | 6.0 × 103 | |
HOLMES | Cas12a | PCR | 88 (PCR) and 15 (CRISPR) | Column-based | Fluorescence | SNP discrimination in cell lines and human samples; detection of viruses (PRV, JEV); virus-strain discrimination | 1.0 × 10–17 | 6.6 × 103 | |
CRISPR-materials | Cas12a | RPA | 40 (RPA) and 240 (CRISPR) | Synthetic targets | Fluorescence or μPAD (visual and electronic) | Detection of EBOV synthetic RNA | 1.0 × 10–17 | 6.6 × 103 | |
CDetection | Cas12b | RPA | 10 (RPA) and 60–180 (CRISPR) | Synthetic targets or crude extraction | Fluorescence | Detection of HPV16; human ABO blood genotyping; BRCA1 and TP53 SNPs | 1.0 × 10–18 | 6.0 × 102 | |
HOLMESv2 | Cas12b | LAMP | 40 (LAMP) and 35 (CRISPR) or 120 (one pot) | NS | Fluorescence | SNP discrimination in cell lines; RNA virus detection (JEV); human mRNA and circular RNA detection; DNA methylation | 1.0 × 10–17 | 6.0 × 103 | |
E-CRISPR | Cas12a | – | 30–180 | Synthetic targets (nucleic acids) | Electrochemical | Detection of viruses (HPV16, PB19) and protein (TGF-ß1) | 5.0 × 10–11 | 3.0 × 1010 | |
CRISPR type VI | |||||||||
– | Cas13 | – | NS | NS | Fluorescence | Detection of human mRNA; detection of bacteriophage λ-RNA | 1.0 × 10–12 | 6.0 × 108 | |
SHERLOCK | Cas13 | NASBA or RPA | 132 (NASBA) or 120 (RPA) and 60–180 (CRISPR) | Column-based or crude extraction | Fluorescence | Detection of viruses (ZIKV, DENV) and bacteria (E. coli, K. pneumoniae, P. aeruginosa, M. tuberculosis, S. aureus); discrimination between virus strains; detection of SNPs | 2.0 × 10–18 | 1.2 × 103 | |
SHERLOCKv2b | Cas13 | RPA | 60 (RPA) and 60–180 (CRISPR) or 60–180 (one pot) | Column-based or crude extraction | Fluorescence or lateral flow | Detection of viruses (ZIKV, DENV) and bacteria (P. aeruginosa, S. aureus); discrimination between virus strains; detection of SNPs | 8.0 × 10–21 | 4.8 | |
SHINEb | Cas13 | RPA | 50 (one pot) | Crude extraction | Fluorescence or lateral flow | Detection of SARS-CoV-2 | 8.3 × 10–18 | 5.0 × 103 | |
STOPCovidb | Cas12b | LAMP | 60 (one pot) | Crude extraction | Fluorescence or lateral flow | Detection of SARS-CoV-2 | 3.3 × 10–18 | 2.0 × 103 | |
CARMEN | Cas13 | PCR or RPA | 20 (RPA) and 180 (CRISPR) | Column-based | Fluorescence | Detection of 169 viruses; subtyping of influenza A strains; detection of HIV drug-resistant mutations | 9 × 10–19 | 5.4 × 102 | |
APC-Cas | Cas13 | Allosteric-probe-initiated amplification with DNA polymerase | 110 (APC) and 30 (CRISPR) | None | Fluorescence | Detection of S. enteritidis | One colony-forming unit | – | |
Cas13 | – | <240 | Column-based | Electrochemical | Detection of microRNAs (miR-19b and miR-20a) | 1 × 10–11 | 6.0 × 109 | ||
PECL-CRISPR | Cas13 | EXPAR | 30 (CRISPR), 30 (phosphorylation of pre-trigger), 30 (EXPAR) | Column-based | Electrochemiluminescence | Detection of microRNAs (miR-17, let‐7 family miRNAs) | 1.0 × 10–15 | 6.0 × 105 | |
- NS, not specified; APC-Cas, allosteric probe-initiated catalysis and CRISPR-Cas13a system; BRCA1, breast cancer 1 gene; circRNA, circular RNA; Cas9nAR, Cas9 nickase-based amplification reaction; CRISDA, CRISPR–Cas9-triggered nicking endonuclease-mediated strand-displacement amplification; DENV, dengue virus; DMD, Duchenne muscular dystrophy; EBOV, Ebola virus; E. coli, Escherichia coli; HERC2, HECT and RLD domain containing E3 ubiquitin protein ligase 2 gene; HPV, human papillomavirus; JEV, Japanese encephalitis virus; K. pneumoniae, Klebsiella pneumoniae; KRAS, KRAS proto-oncogene GTPase; M. smegmatis, Mycobacterium smegmatis; M. tuberculosis, Mycobacterium tuberculosis; PECL, portable electrochemiluminescence chip; P. aeruginosa, Pseudomonas aeruginosa; PB19, parvovirus B19; PRV, pseudorabies virus; S. erythraea, Saccharopolyspora erythraea; S. aureus, Staphylococcus aureus; S. enteritidis, Salmonella enteritidis; S. typhimurium, Salmonella typhimurium; TP53, tumour protein P53 gene.
- aAssay time indicates the approximate incubation time most frequently used in the referred study (different assay times can be reported, depending on the intended sensitivity and the readout).
- bPOC compatibility indicates whether the entire assay as reported—including sample preparation (that is, crude extraction) and readout—can be performed at POC or in the field with minimal equipment.
- cLimits of detection cannot always be directly compared across studies, in particular because some studies did not report how the LOD was determined, or reported the target concentration either in the transport media of the sample or in the final reaction. The LODs shown here reflect the optimal LODs reported. In general, LODs depend on the type of input material (raw or synthetic), type of readout and incubation time.